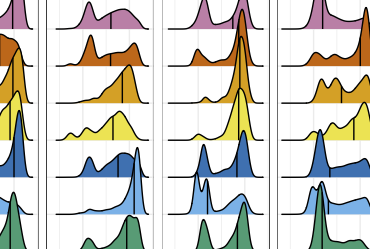

The Human Microbiome Compendium: An ongoing project to build a dataset of the human microbiome at an unprecedented scale
Integration of 168,000 samples reveals global patterns of the human gut microbiome (Cell, 2025)
Full dataset for download, version 1.1 (Zenodo)
MicroBioMap R package (github)
Website (microbiomap.org)
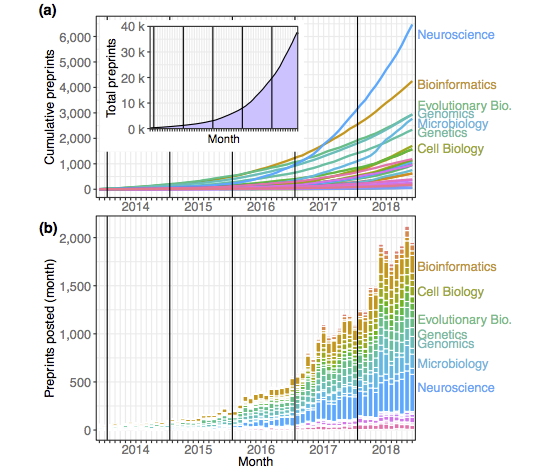

A website to sort through bioRxiv preprints and find relevant, trending papers
Rxivist.org website
Data & code (GitHub)
API documentation
Our paper on preprint trends using Rxivist data (eLife 2019)
Our paper describing the Rxivist.org website (PLOS Biology 2019)
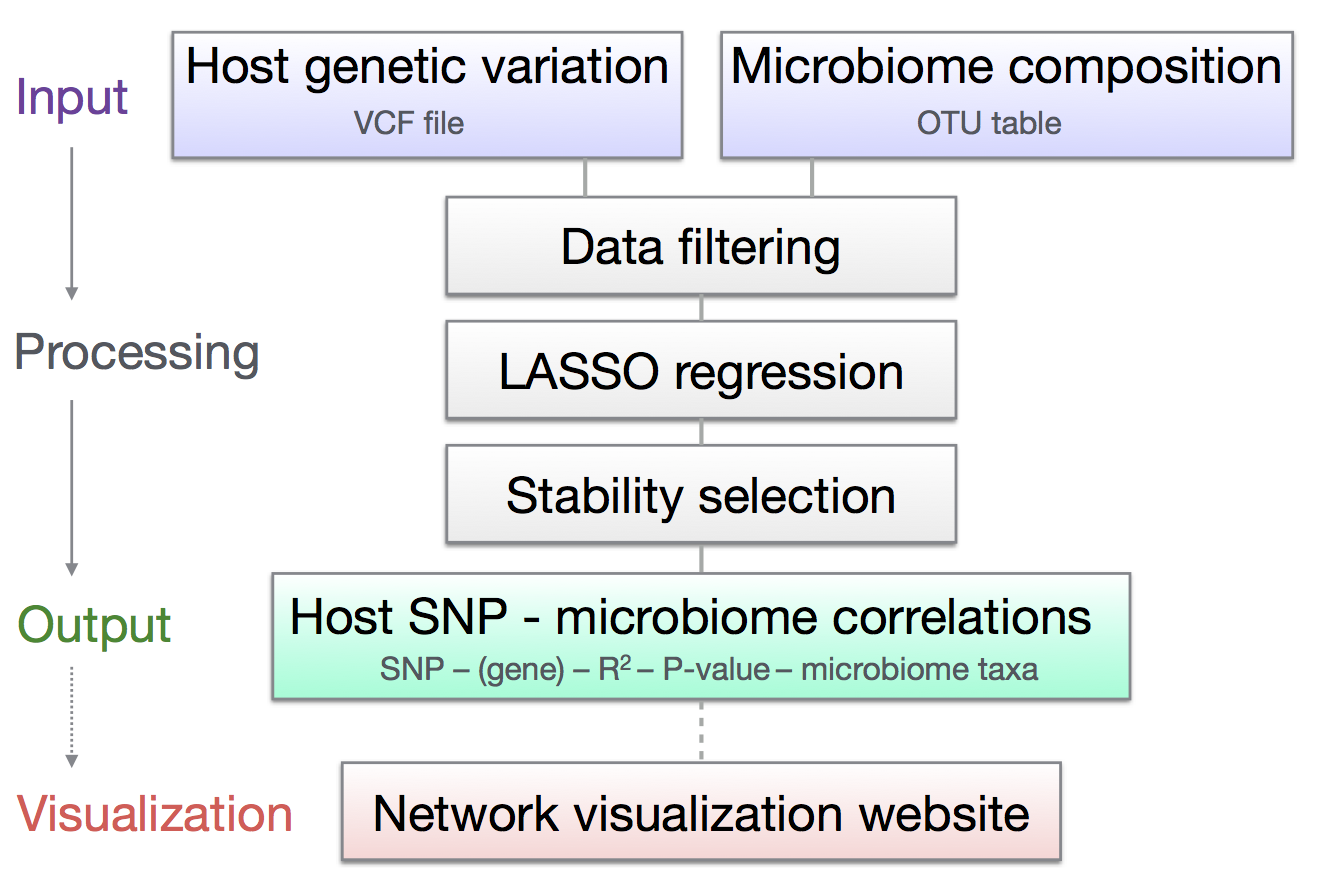

A program to identify associations between host genetic variation and microbiome composition using machine-learning
Software, code, installation instructions, and tutorial (GitHub)
Online network visualization tool
Paper (GigaScience)
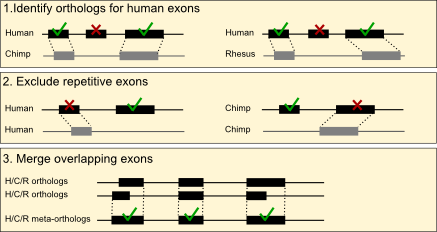
Primate Orthologous Exon Database
This database provides a catalog of unique, non-overlapping, orthologous exon regions in the genomes of human, chimpanzee, and rhesus macaque. The database can be used in analysis of multi-species RNA-seq data, allowing for comparisons of exon-level expression across primates, as well as comparative examination of alternative splicing and transcript isoforms.
Note: the Primate Orthologous Exon Database is out of date and no longer being updated or supported.
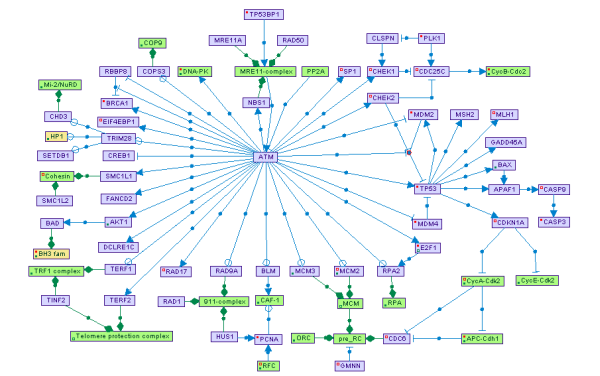
SPIKE
SPIKE is a database of highly curated human signaling pathways with an associated interactive software tool. Users can view and download individual pathway maps and browse the entire database from this website, or install a stand-alone software tool that allows dynamic visualization and manipulation of the database and additional analyses.
Data from Morton et al. (2015) PLOS Genetics
All Supplementary Tables and FiguresFigshare
All raw sequence data (fastq files)
MG-RAST
Data from Burns et al. (2014) Genome Medicine
All Supplementary Data files, including OTU, pathway, and enzyme abundance tablesGenome Medicine
Data from Blekhman et al. (2010) Genome Research
Supplementary Table 1Tab-delimited text file (zip)
Data from Blekhman et al. (2008) PLoS Genetics
Supplementary dataTab-delimited text file (zip)
hOMIM data from Blekhman et al. (2008) Current Biology
A curated dataset of human mendelian disease genes.Tab-delimited text file

